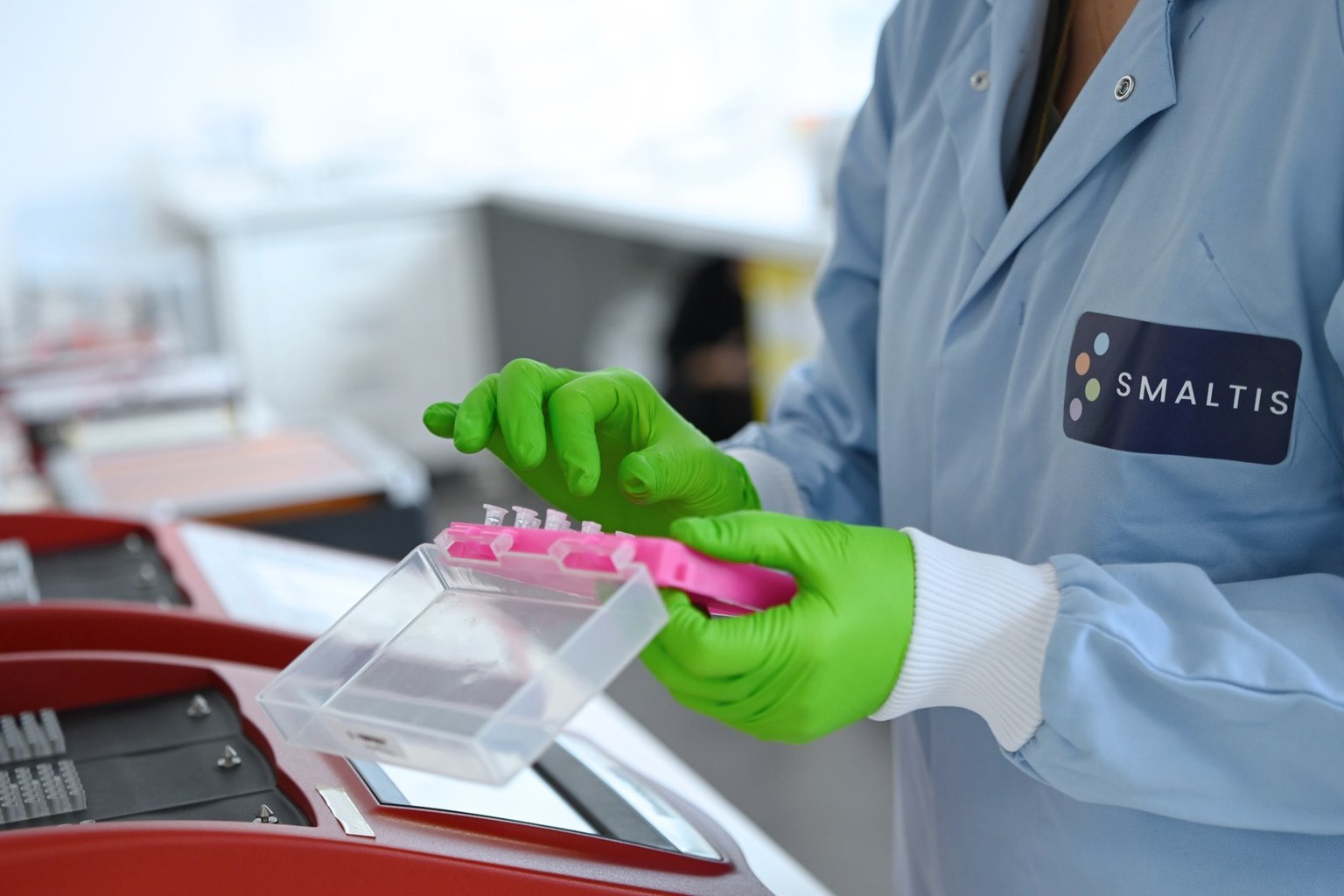

Starting point: Whole Genome Sequencing
In current practice, Whole Genome Sequencing (WGS) has become the foundation of candidate strain characterization. It makes it possible to confirm taxonomic identity, detect genes associated with antimicrobial resistance, virulence, or other safety determinants, and examine the genetic environment of these sequences: plasmids, mobile elements, integration regions, or other supports likely to promote transfer. [1-3]
However, the presence of a gene is not, by itself, proof of a functional risk. A putative determinant may be truncated, silent, not translated, or non-functional. Conversely, a hit on a clinically relevant AMR gene remains a signal that must be investigated seriously. The correct interpretation is therefore to cross four levels of information: presence of the determinant, sequence integrity, genetic context, and consistency with the phenotype. This is where one moves from simple genomic annotation to true strain characterization. [2,3]
Antimicrobial resistance: the genome guides, the phenotype decides
A genomic assessment alone is not sufficient. To properly characterize antimicrobial resistance, the phenotype must be measured, in particular the MICs of antibiotics of interest, using methods adapted to the strain under study. For lactic acid bacteria and bifidobacteria, reference frameworks rely in particular on ISO 10932/IDF 223. For other strict anaerobes, the approach is classically based on CLSI M11. At Smaltis, this logic is complemented, when necessary, by a reasoned adaptation of culture conditions and expert interpretation for demanding or atypical strains.
When cut-offs or interpretive thresholds exist, they structure the analysis. When they do not – which is common for new or poorly documented strains – the evaluation has to go further: compare with control strains, reason in terms of wild-type versus non-wild-type profiles, use phylogenetically relevant comparators, and relate the results to the genetic context observed by WGS. The challenge is not simply to say that a strain is ‘susceptible’ or ‘resistant’ to a given molecule, but to understand whether the observed signal reflects an intrinsic background, an acquired determinant, or a potentially transferable mechanism. [2,3]

Safety: going beyond genomic keywords
The safety of a probiotic strain cannot be reduced to its levels of resistance to antibiotics. The search for genes associated with virulence, toxigenicity, or other undesirable determinants is essential, but here again it does not replace targeted experimental characterization. Depending on the strain and the intended application, this may lead to documenting the absence of hemolysis, the absence of cytotoxic effects, the absence of deleterious impact on epithelial integrity, or other signals compatible with an opportunistic profile. The objective is to generate a level of evidence that is consistent with the strain’s actual risk. [1,2,7]
In other words, the right strategy is to start from the WGS data, identify points of attention, and then select the in vitro assays that will confirm, rule out, or clarify those signals. [1-3]
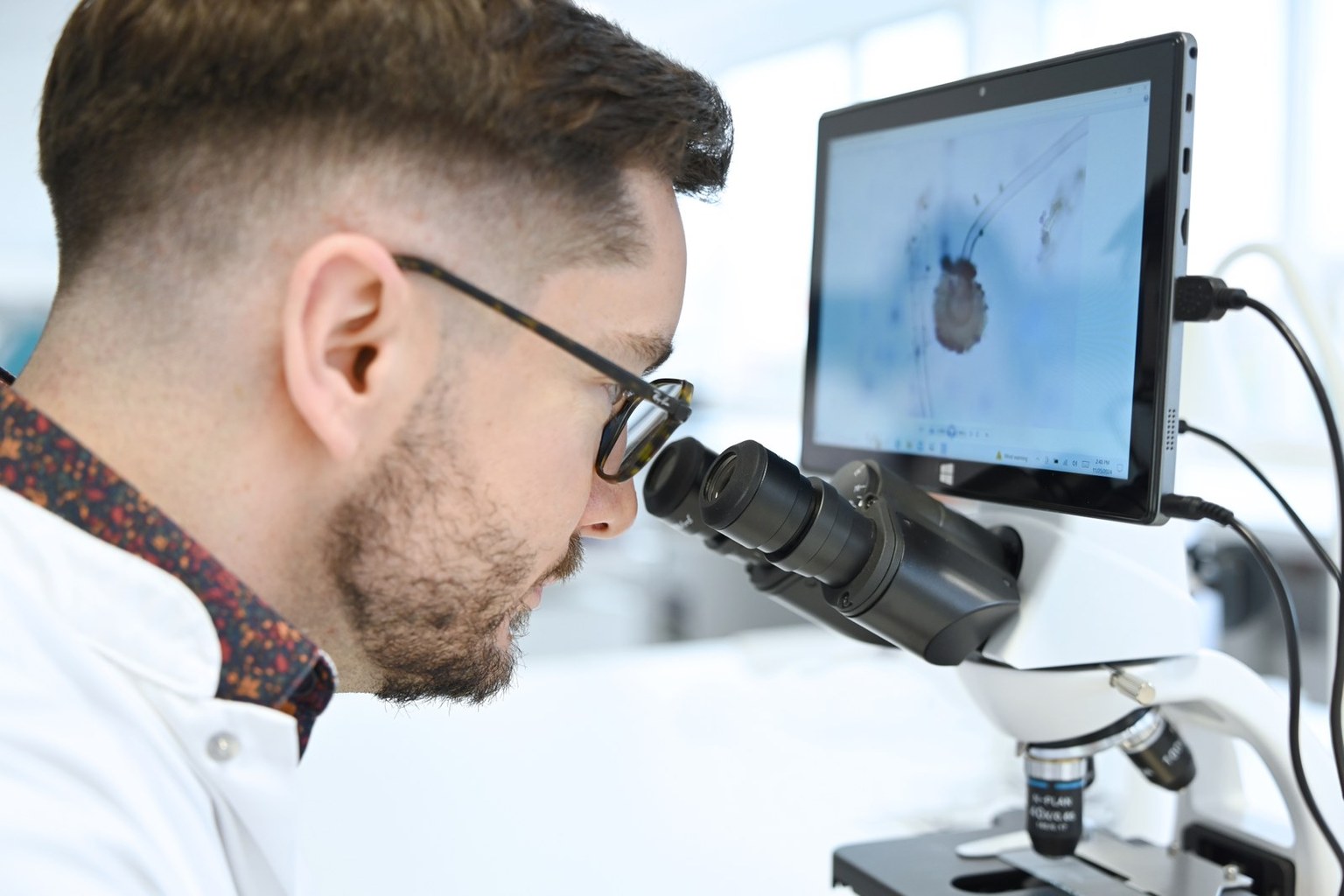

Efficacy: what in vitro models can demonstrate, and what they cannot
The objective is to produce mechanistic and discriminating evidence that is useful for selecting, comparing, and understanding strains. In vitro assays can therefore support the biological plausibility of a benefit, but they do not replace clinical demonstration when a health claim is to be made in the strict sense. [5,7,8]
Three areas are particularly relevant. The first concerns interactions with the microbiota and pathogens: growth inhibition, production of antagonistic molecules, competition for adhesion, or exclusion of undesirable bacteria. The second concerns the intestinal barrier, with models such as Caco-2, HT-29, or T84 used to assess TEER, permeability, tight junctions, or the mucosal response. The third concerns immunomodulation, through measurable signals such as cytokines and the responses of epithelial cells, macrophages, or PBMCs. In every case, the value of in vitro models lies in documenting a plausible mechanism of action and ranking strains, not in claiming a clinical benefit on their own. [9-11]
The gut-brain axis is a case apart. It is a very attractive field, but one that is still only weakly standardized in the laboratory. Conventional 2D models capture only part of it indirectly. The most relevant approaches today are integrated systems such as organoids and gut-on-chip or even gut-brain-on-chip devices. These models are promising, but they remain advanced tools rather than universal routine standards. [12]

 All articles
All articles